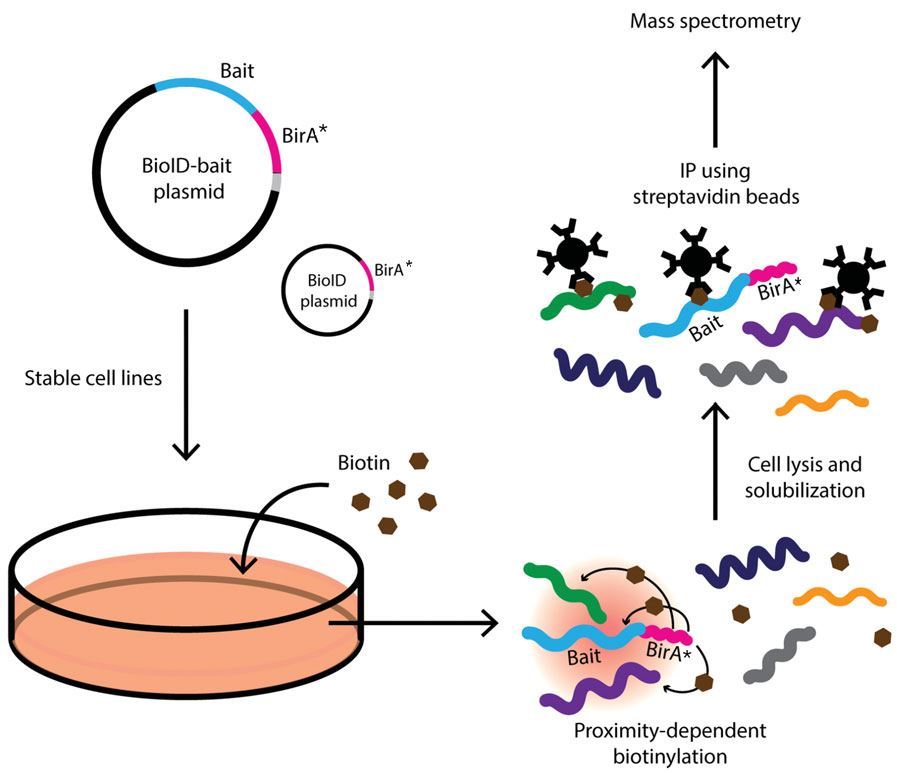
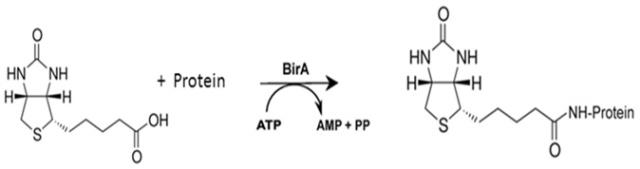
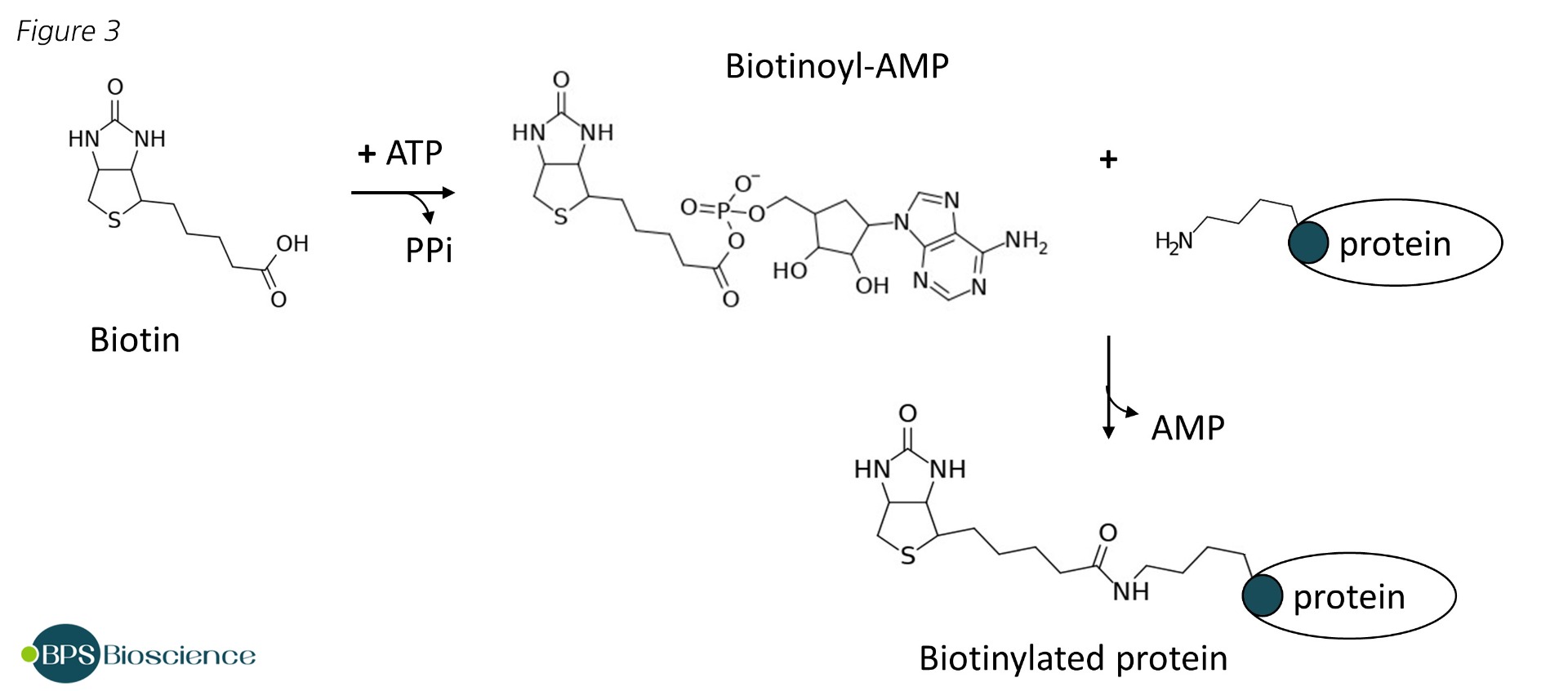
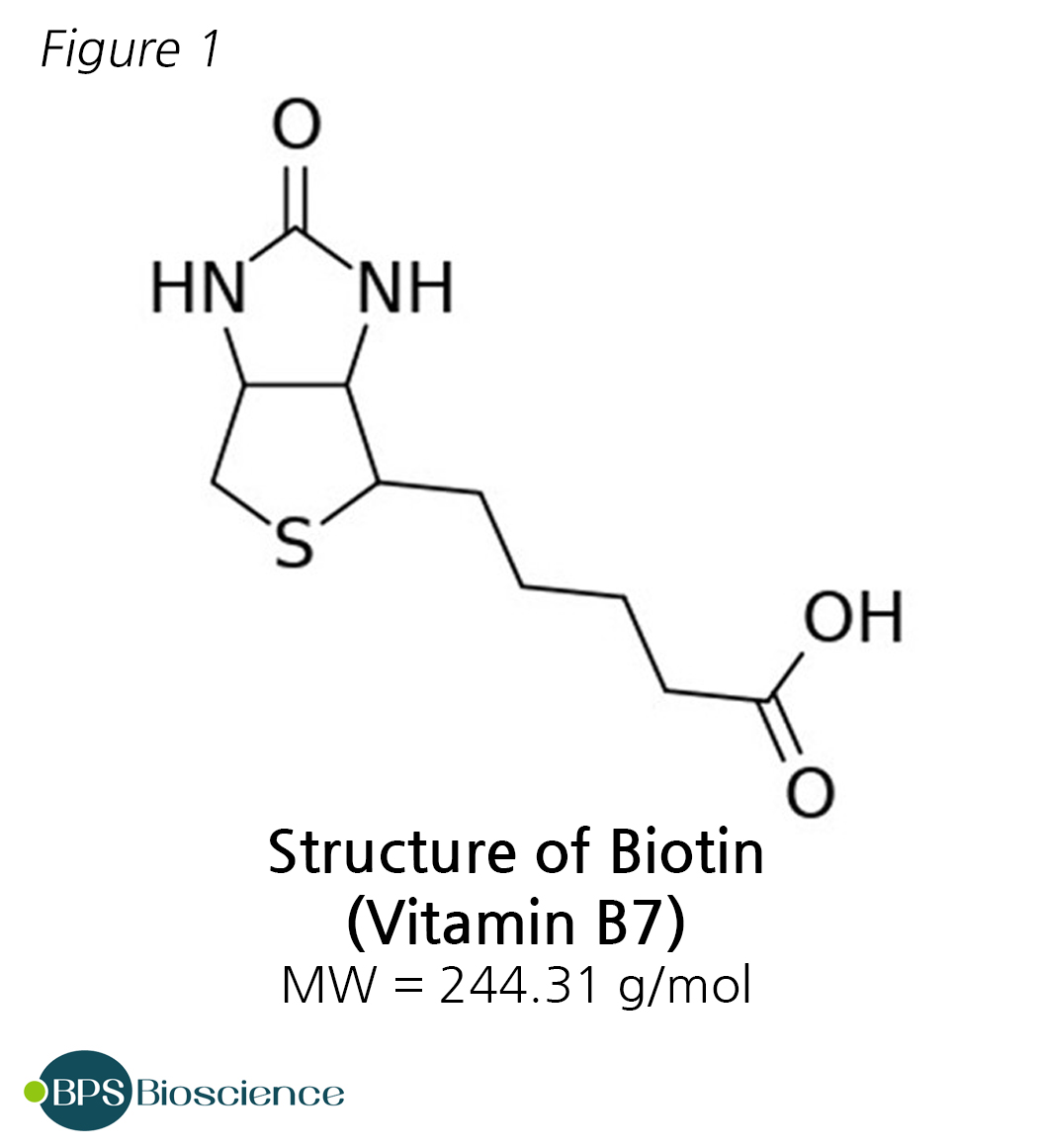
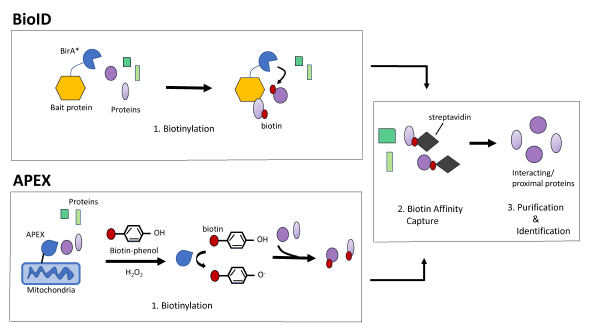
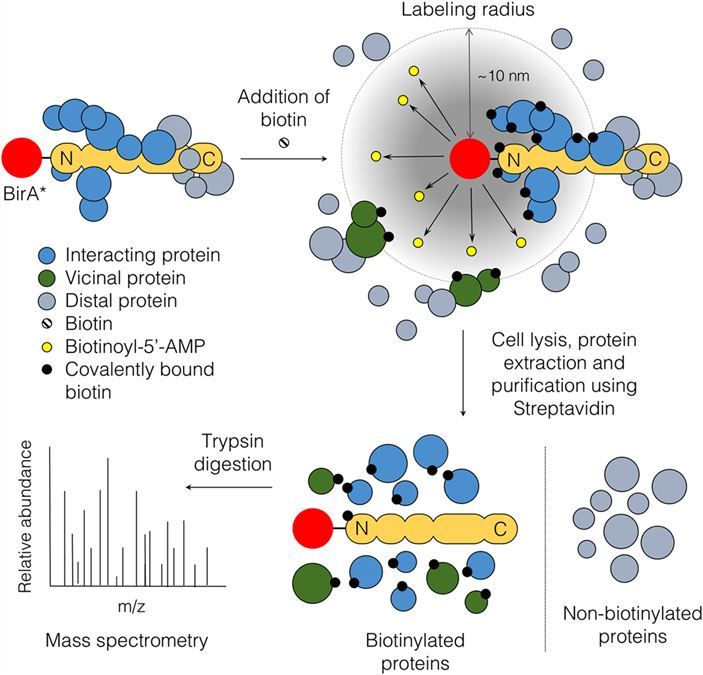
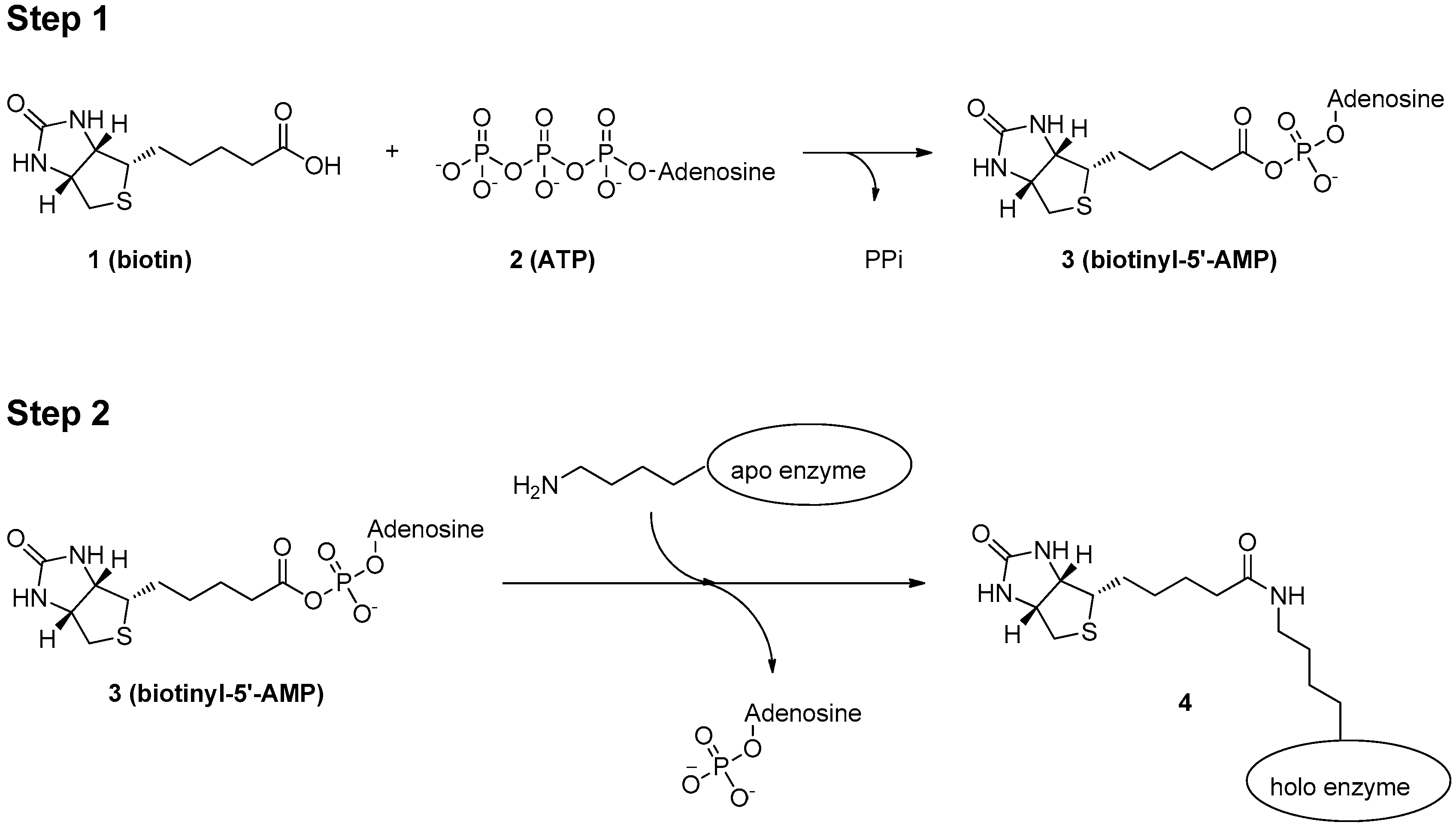
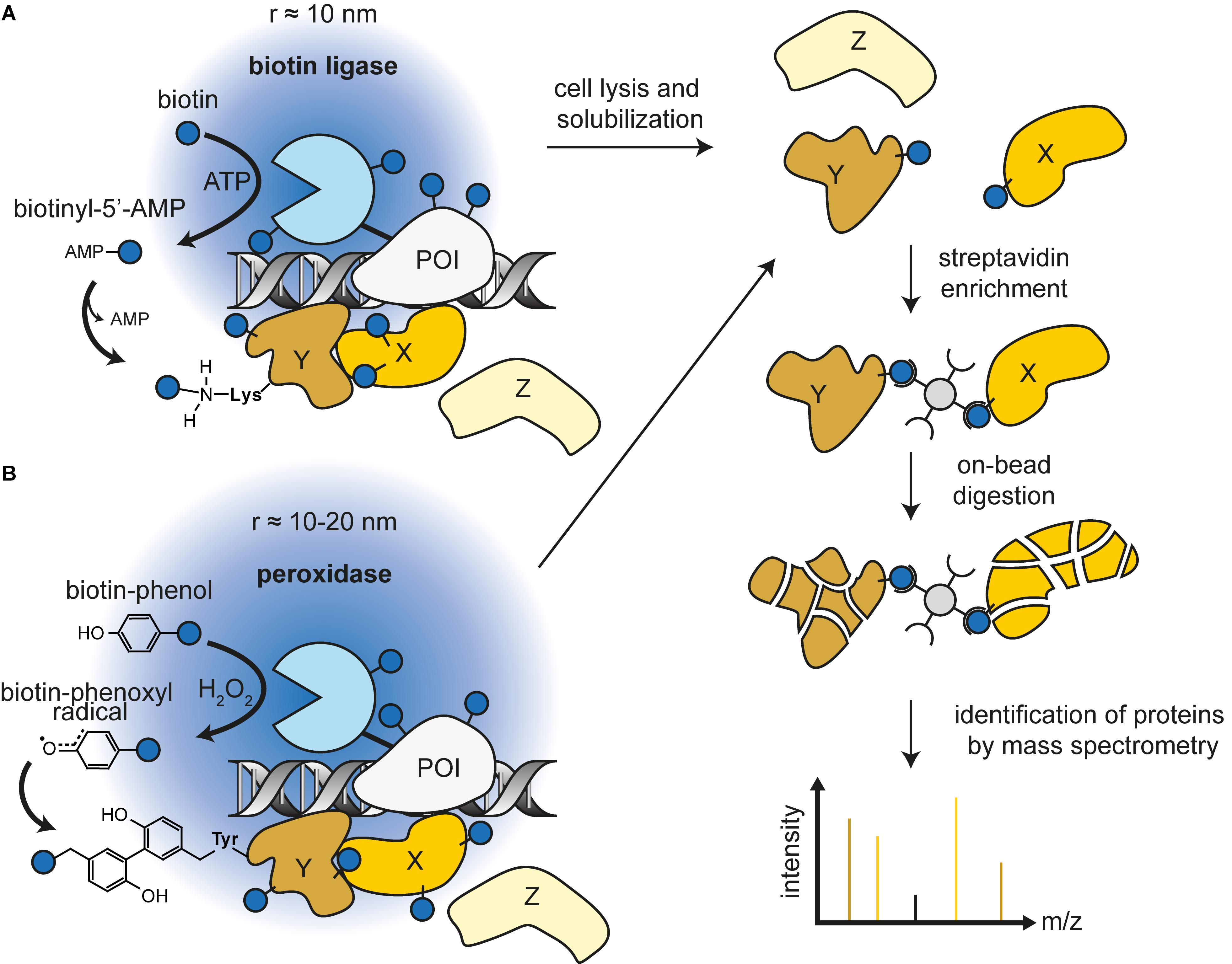
Identification of Interactions in the NMD Complex Using Proximity-Dependent Biotinylation (BioID) | PLOS ONE

Site-specific labeling of cell surface proteins with biophysical probes using biotin ligase | Nature Methods
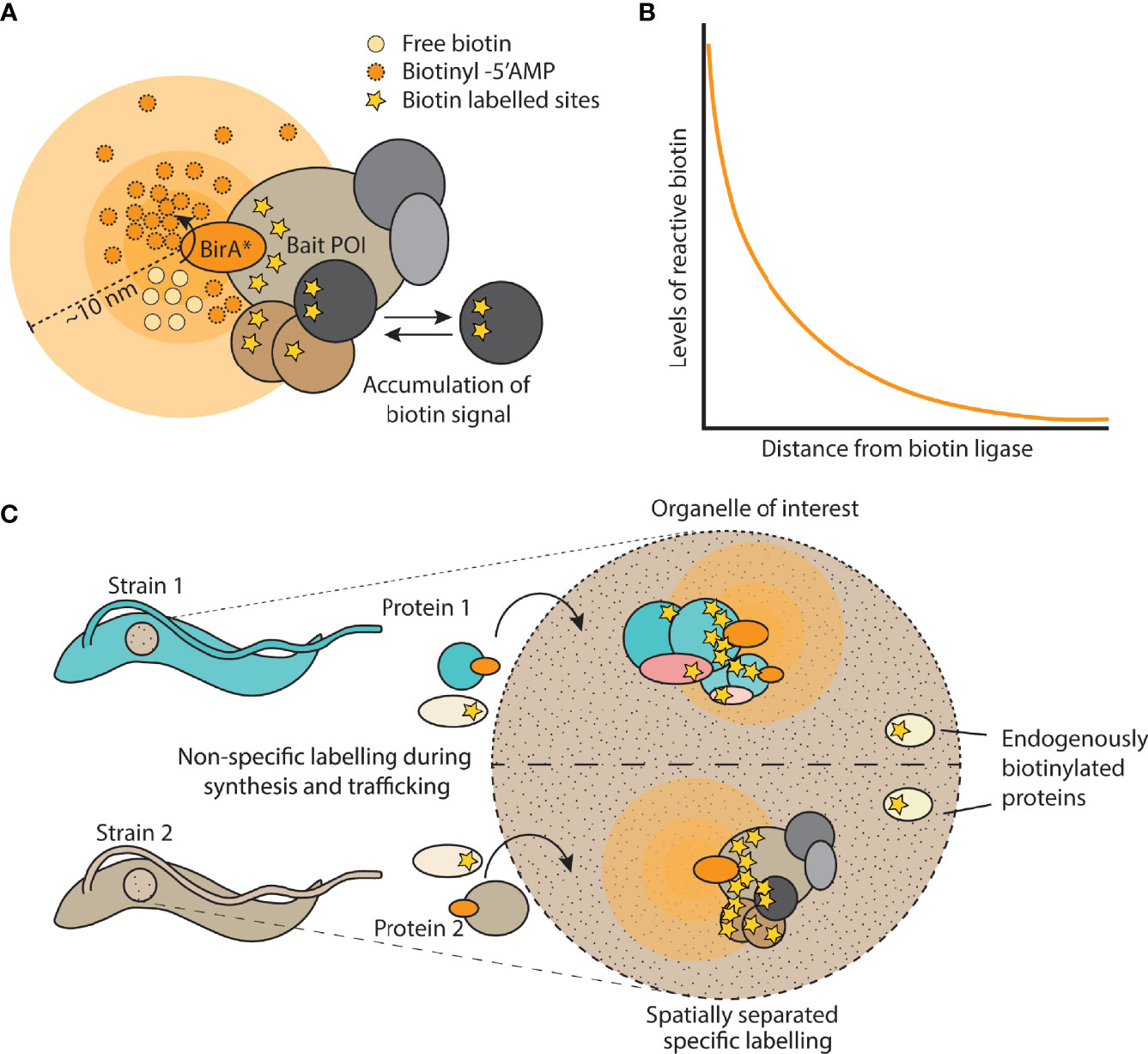
Frontiers | Tag Thy Neighbour: Nanometre-Scale Insights Into Kinetoplastid Parasites With Proximity Dependent Biotinylation
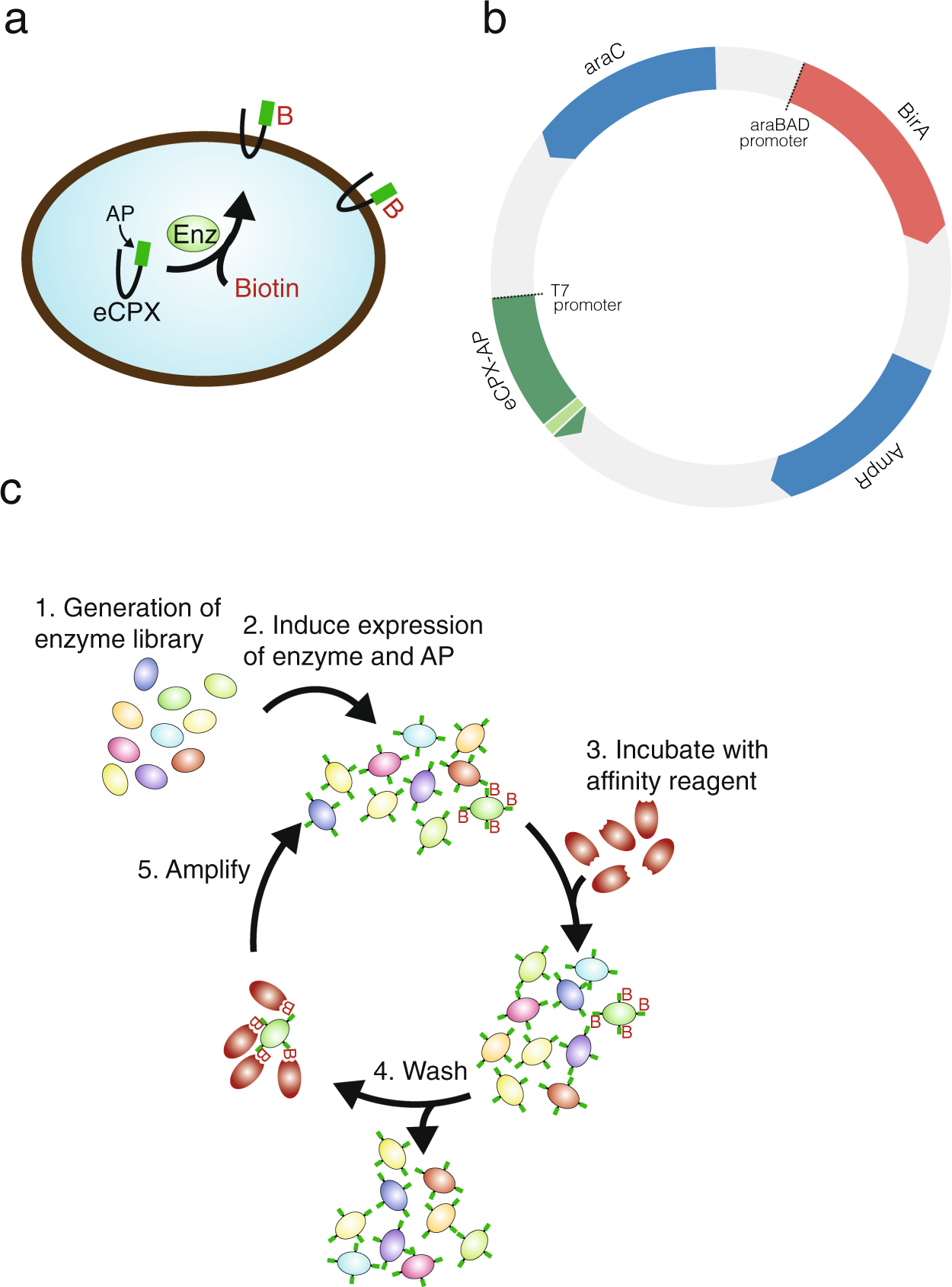
A bacterial display system for effective selection of protein-biotin ligase BirA variants with novel peptide specificity | Scientific Reports
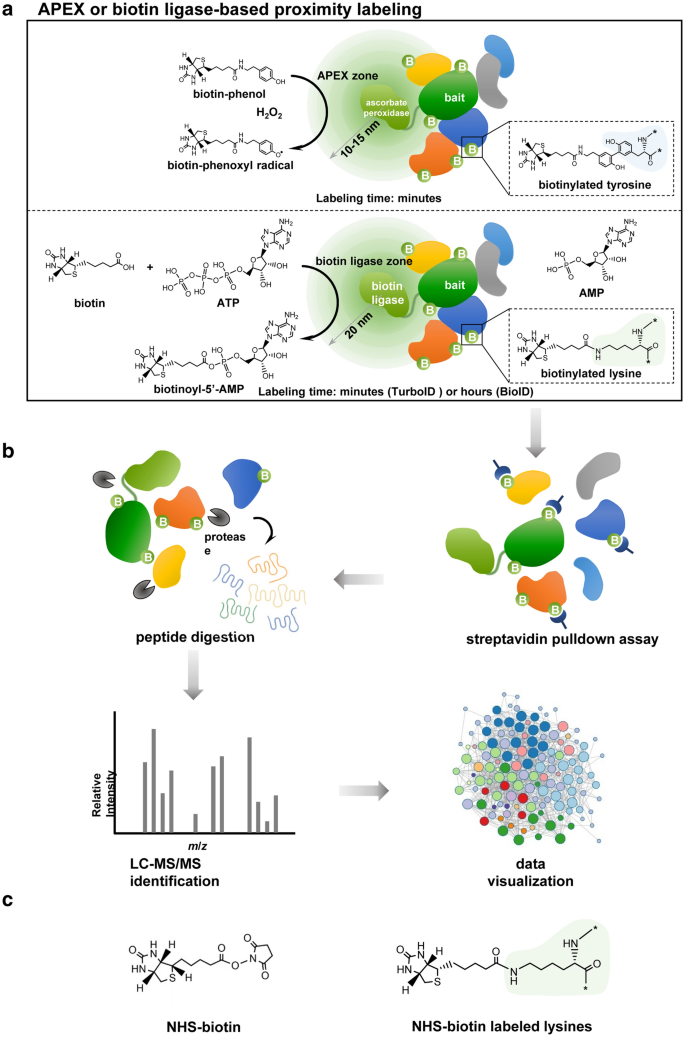
In vivo interactome profiling by enzyme‐catalyzed proximity labeling | Cell & Bioscience | Full Text
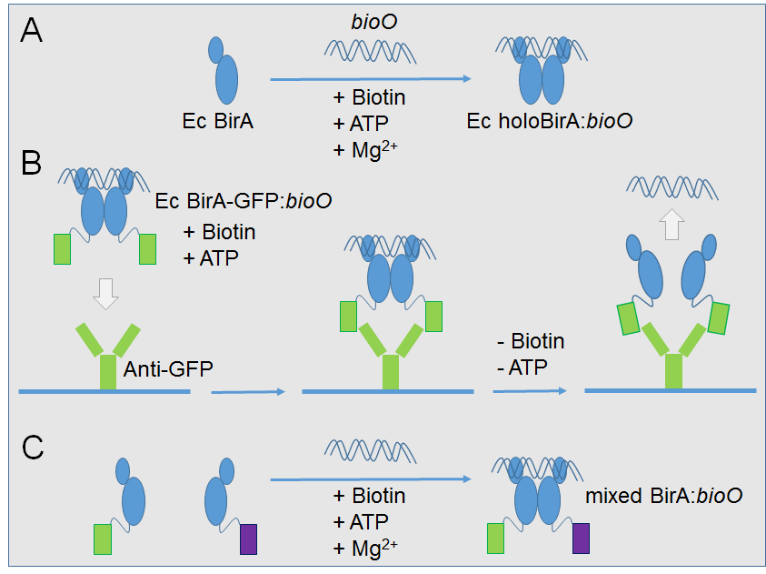
Biotin Protein Ligase as Protein: DNA Connecting System - Research and Innovation Services - JCU Australia

Sulfonamide-Based Inhibitors of Biotin Protein Ligase as New Antibiotic Leads | ACS Chemical Biology

The Atypical Occurrence of Two Biotin Protein Ligases in Francisella novicida Is Due to Distinct Roles in Virulence and Biotin Metabolism | mBio

Figure 1 from A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells | Semantic Scholar
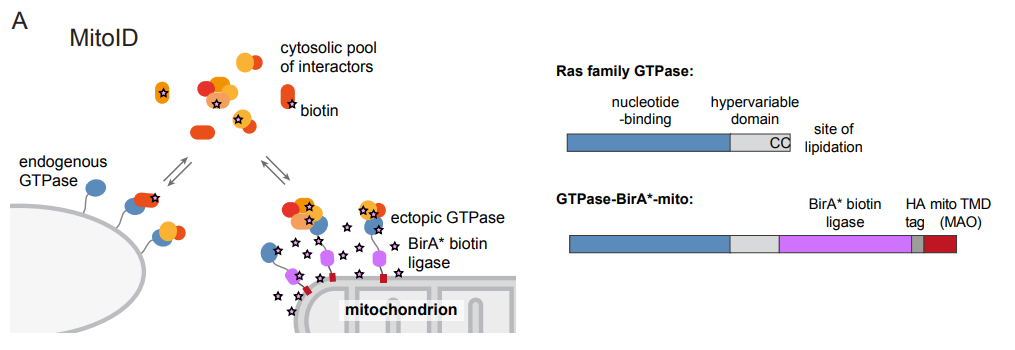
In vivo identification of GTPase interactors by mitochondrial relocalization and proximity biotinylation - preLights